Jflow
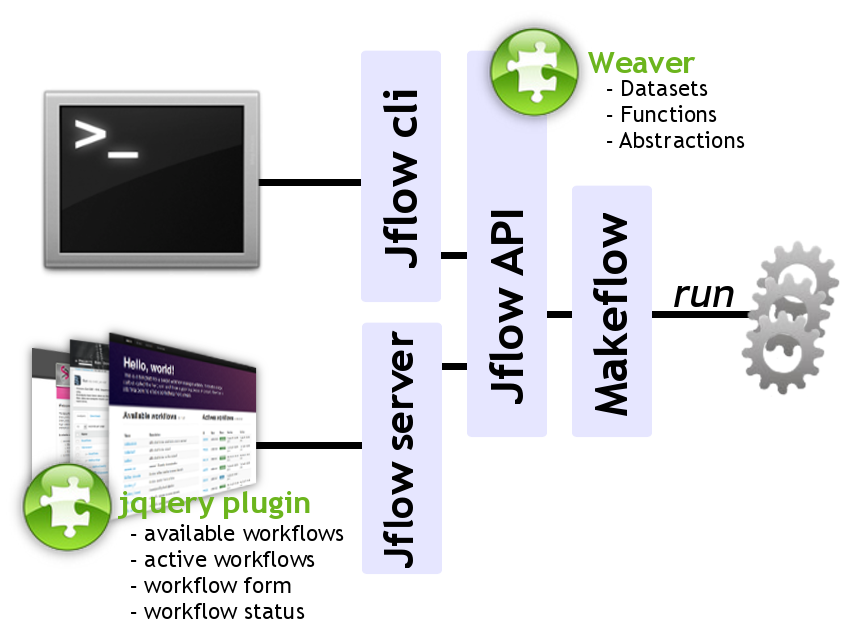
A JavaScript based workflow management system, composed of Jquery plugins which can easily be embedded in any WEB application and a Python library providing all requested features to setup, run and monitor workflows.
Jérôme Mariette, Frédéric Escudié, Philippe Bardou, Ibouniyamine Nabihoudine, Céline Noirot, Marie-Stéphane Trotard, Christine Gaspin, Christophe Klopp. Jflow: a workflow management system for web applications. Bioinformatics. 2015. doi 10.1093/bioinformatics/btv589. - Abstract/FREE Full Text
Jflow Summary
Today biologists produce large data sets and are in demand of rich and simple to use WEB portals in which they can upload and analyse their files. Providing such tools requires to mask the complexity induced by the needed High Perfomance Computing (HPC) environment. The connexion between interface and computing infrastructure is usually specific to each portal. We introduce jflow, a Workflow Management System (WMS), composed of Jquery plugins which can easily be embedded in any WEB application and a Python library providing all requested features to setup, run and monitor workflows.
Jflow Availability
Jflow is available under the GNU General Public License (GPL). The package is comming with a full documentation a quickstart and a running test portal. Sources are available here.
Plugin insertion
Learn how to insert the jflow plugin within your own application.
Add new workflows
Learn how to add new workflows.
Add new components
Your workflow requires new component, learn how to add new ones.
Example
Check out an example of jflow integration.